Sept. 19, 2022
Contact: Eric Stann, 573-882-3346, StannE@missouri.edu
There was once a time, not so long ago, when scientists like Casey Holliday needed scalpels, scissors and even their own hands to conduct anatomical research. But now, with recent advances in technology, Holliday and his colleagues at the University of Missouri are using artificial intelligence (AI) to see inside an animal or a person — down to a single muscle fiber — without ever making a cut.
Holliday, an associate professor of pathology and anatomical sciences, said his lab in the MU School of Medicine is one of only a handful of labs in the world currently using this high-tech approach.
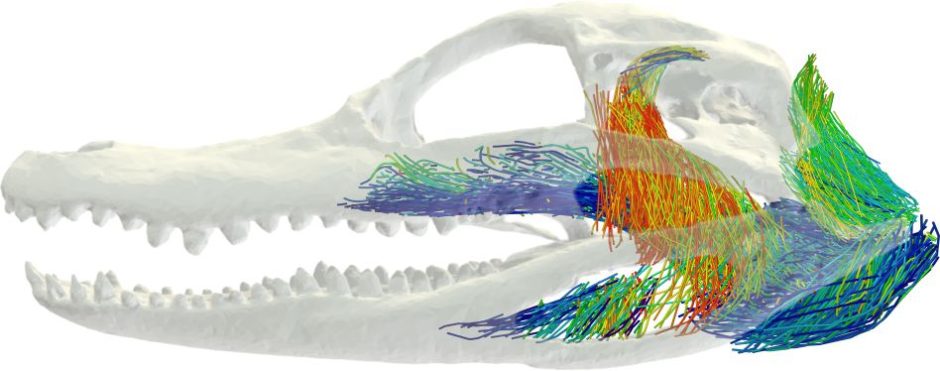
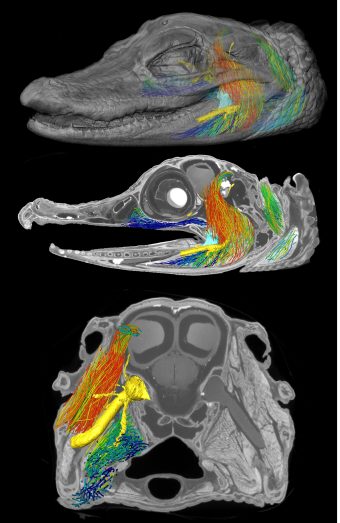
AI can teach computer programs to identify a muscle fiber in an image, such as a CAT scan. Then, researchers can use that data to develop detailed 3D computer models of muscles to better understand how they work together in the body for motor control, Holliday said.
Holliday, along with some of his current and former students, did that recently when they began to study the bite force of a crocodile.
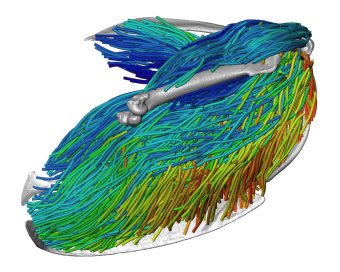

“The unique thing about crocodile heads is that they are flat, and most animals that have evolved to bite really hard, like hyenas, lions, T. rexes and even humans have really tall skulls, because all those jaw muscles are oriented vertically,” Holliday said. “They’re designed that way so they put a big vertical bite force into whatever they're eating. But a crocodile’s muscles are oriented more horizontally.”
The 3D models of muscle architecture could help the team determine how muscles are oriented in crocodile heads to help increase their bite force. Helping to lead this effort is one of Holliday’s former students, Kaleb Sellers, who is now a postdoctoral researcher at the University of Chicago.
“Jaw muscles have long been studied in mammals with the assumption that relatively simple descriptors of muscle anatomy can tell you a great deal about skull function,” Sellers said. “This study shows how complex jaw muscle anatomy is in a reptile group.”
Holliday’s lab first began experimenting with 3D imaging several years ago. Some of their early findings were published in 2019 with a study in Integrative Organismal Biology that showed the development of a 3D model of the skeletal muscles in a European starling.
Transitioning into a digital world
Historically, Holliday said anatomical research — and much of what he did growing up — involved dissecting animals with a scalpel or scissors, or what he calls an “analog” approach. He was first introduced to the benefits of using digital imaging to study anatomy when he joined the “Sue the T. rex” project in the late 1990s. To date, it remains one of the largest and most well-preserved Tyrannosaurus rex specimens ever discovered.
Holliday recalls the moment when the T. rex’s giant skull was transported to Boeing’s Santa Susana Field Laboratory in California to be imaged in one of the aerospace company’s massive CAT scanners normally used to scan jet engines on commercial airplanes.
“At the time, it was the only CAT scanner in the world big enough to fit a T. rex skull, and also had the power needed to push X-rays through rocks,” Holliday said. “Coming out of college I had looked at becoming a radiology technician, but with the Sue project I was learning all about how they CAT scanned this thing, and that really caught my fancy.”
Nowadays, Holliday said many of his current and former students at MU are learning to understand anatomy by using the “cutting edge” imaging and modeling methods that he and his colleagues are creating. One of those students is Emily Lessner, a recent MU alumna who developed her passion for “long-dead animals” by working in Holliday’s lab.
“The digitization process is not only useful to our lab and research,” Lessner said. “It makes our work shareable with other researchers to help hasten scientific advancement, and we can also share them with the public as educational and conservation tools. Specifically, my work looking at the soft tissues and bony correlates in these animals has not only created hundreds of future questions to answer, but also revealed many unknowns. In that way, not only did I gain imaging skills to help with my future work, but I now have more than a career-worth of avenues to explore.”
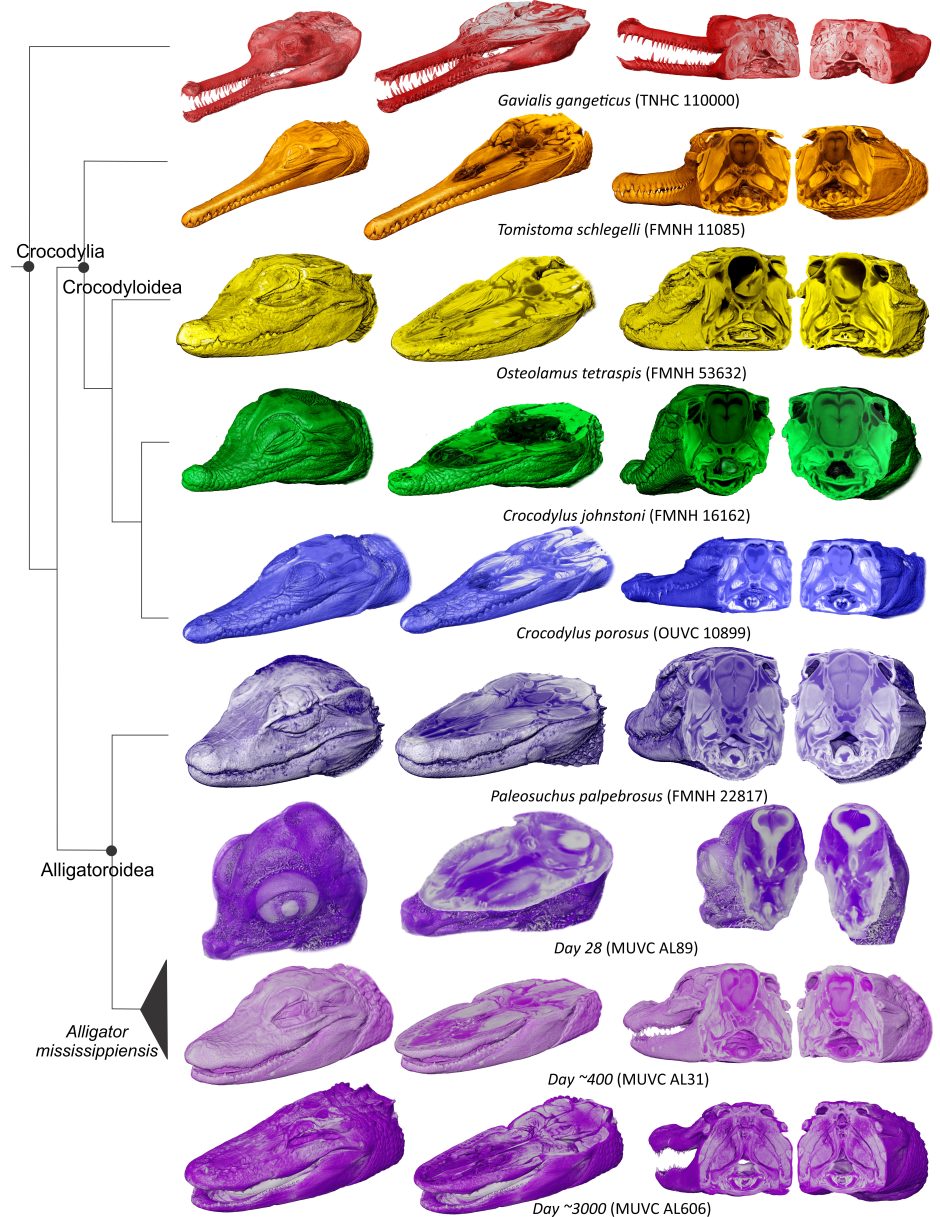

Holliday said plans are also in the works to take their 3D anatomical models a step further by studying how human hands have evolved from their evolutionary ancestors. The project, which is still in its early stages, recently received a grant from the Leakey Foundation. Joining Holliday on the project will be two of his colleagues at MU, Carol Ward, a Curators Distinguished Professor of pathology and anatomical sciences, and Kevin Middleton, an associate professor of biological sciences.
While about 90% of the research done in Holliday’s lab involves studying things that exist in the modern world, he said the data they collect can also inform the fossil record, like additional knowledge about how the T. rex moved and functioned.
“With better knowledge of actual muscle anatomy, we can really figure out how the T. rex could really do fine motor controls, and more nuanced behaviors, such as bite force and feeding behavior,” Holliday said.
Editor’s note:
“New frontiers in imaging, anatomy, and mechanics of crocodylian jaw muscles,” was published in The Anatomical Record. Other authors include Kaleb Sellers, Emily Lessner, Kevin Middleton and Conner Verhulst at MU, Corrine Cranor at South Dakota School of Mines and Technology, Stephan Lautenschlager at University of Birmingham and Matthew Brown and Matthew Colbert at University of Texas-Austin. Funding was provided by grants from the National Science Foundation (EAR/SEB 1631684, NSF IOS PMB 1457319, EAR-1762458 and DBI-1902242), Missouri Research Board, University of Missouri Research Council and Jackson School of Geosciences Geology Foundation. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies.